CSE 599 · Academic Case Study
FNO-Diffusion for Brain MRI Segmentation
I tested a tempting fusion idea: if Fourier Neural Operators can capture global spatial structure, could they improve a diffusion-based brain MRI segmentation pipeline? The answer was no. The diffusion U-shape baseline won, and the failed hybrid became the most useful lesson: dense medical segmentation needs local boundary refinement, not only global context.
The project compared three tracks on BraTS 2021: an FNO segmentation baseline, a diffusion model with a U-shape backbone, and a proposed FNO-Diffusion hybrid. The hybrid underperformed both baselines, which pushed the analysis toward why FNO may belong as a global branch rather than a full replacement for local segmentation pathways.
The Segmentation Problem
The task was not just to detect a tumor. It was to preserve pixel-level structure across multiple tumor subregions.
Dataset
BraTS 2021, 1,251 3D MRI samples
Input
4 modalities: T1, T1Gd, T2, FLAIR
Output
Background, NCR, ED, ET masks
Metric
DICE coefficient
Split
70% train / 20% val / 10% test
Compute
1× NVIDIA Tesla V100, ~2-8 hours

Why the hypothesis sounded reasonable
- Diffusion models can regularize mask generation by learning a denoising process.
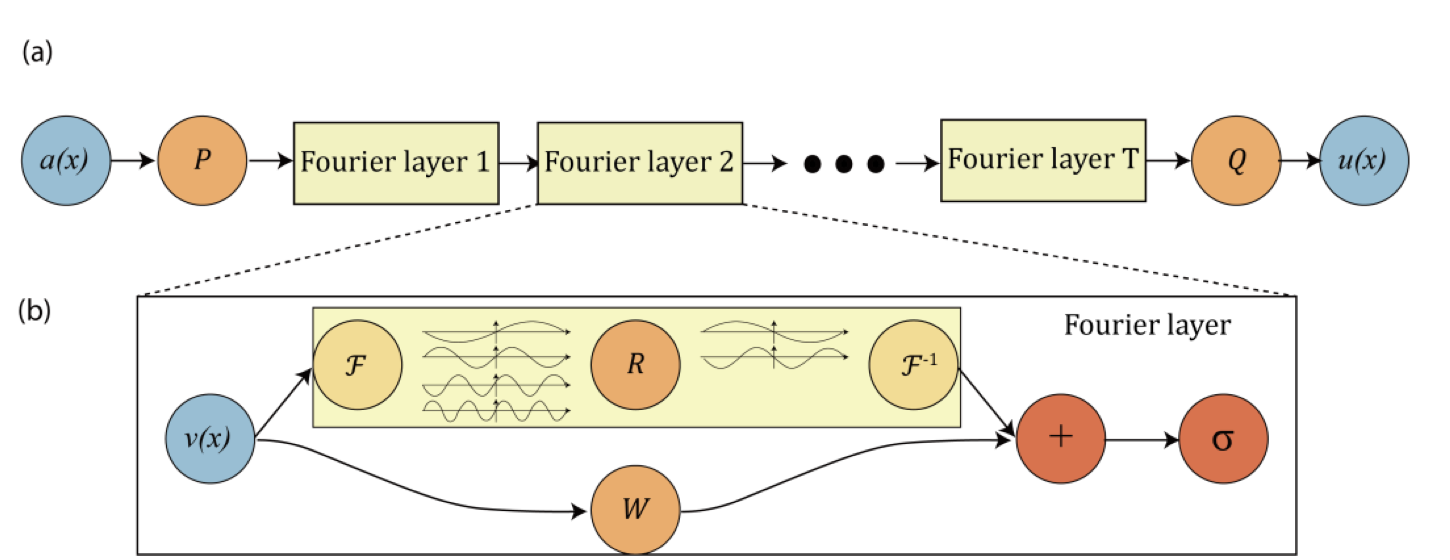
- FNOs model global spatial relationships through spectral convolution.
- Brain tumors are spatial structures, so a global operator seemed like a natural complement to diffusion.
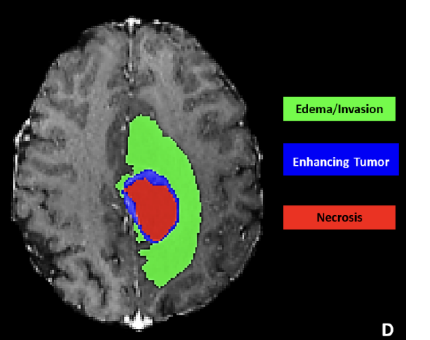
Gray area
Necrotic tumor core
Peritumoral edema
Enhancing tumor
What Each Model Tested
I framed the experiment as three diagnostic questions instead of three disconnected architectures.
A) FNO Segmentation Baseline
Question: Can spectral global modeling segment tumor masks directly?
Result: Competitive but weaker than the U-shape diffusion baseline.
B) Diffusion with U-shape Baseline
Question: Does diffusion-guided supervision plus a local-detail-preserving backbone work better?
Result: Best result, with 69.44% DICE.
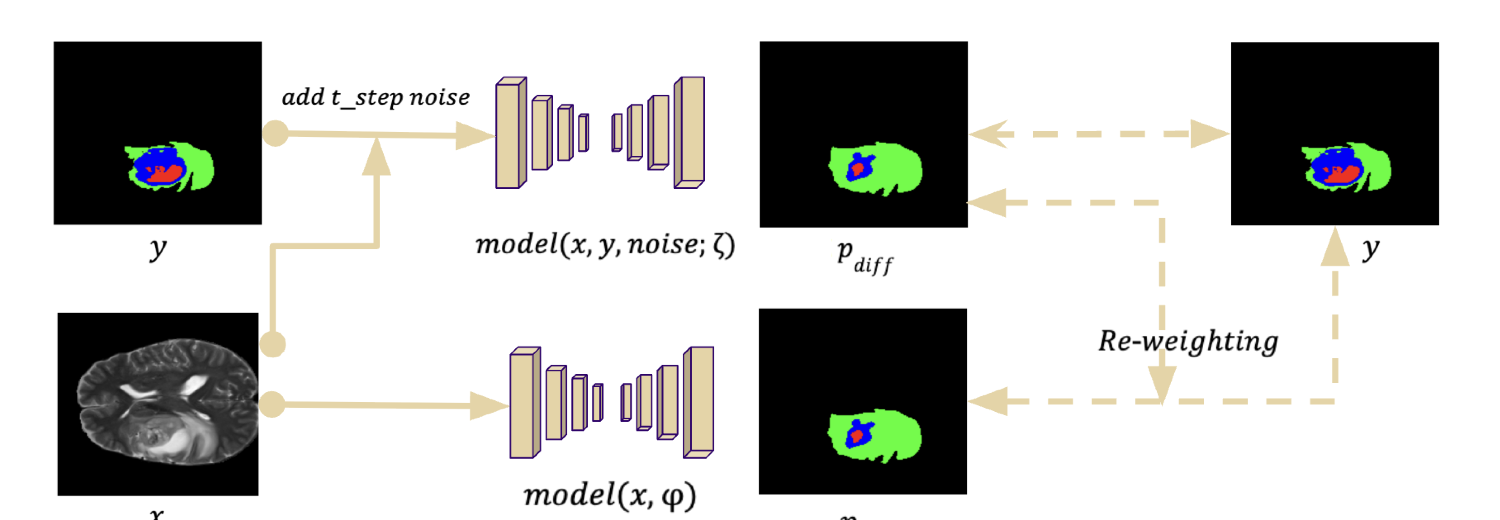
C) Proposed FNO-Diffusion Hybrid
Question: Can FNO improve the diffusion pipeline if inserted into both diffusion and supervised branches?
Result: No. The hybrid fell to 57.03% DICE.
Training Details (optional)
FNO: modes k1=k2=10, width 16, 3 repeated blocks per branch, batch 8, epochs 50, Adam lr 3e-4, GELU, Dice + CE (lambda=0.5).
Diffusion U-shape: Adam lr 1e-2, batch 32, max 300 epochs, early stop 50, EMA 0.99, Dice + CE, dynamic class weights, unsupervised weight 10.
FNO-Diffusion: SGD momentum 0.9, weight decay 3e-5, lr 0.001, batch 32, EMA 0.99, timesteps 1000 (sampling 10), FNO modes [16,16], width 32, blocks/channel 3, time embedding 512, dropout 0.5.
The Result: The Simple U-shape Won
The hybrid did not validate the original hypothesis. It made the segmentation worse.
| Rank | Model | DICE |
|---|---|---|
| 1 | Diffusion with U-shape | 69.44% |
| 2 | FNO baseline | 65.13% |
| 3 | FNO-Diffusion hybrid | 57.03% |
#1 Diffusion with U-shape
DICE: 69.44%
#2 FNO baseline
DICE: 65.13%
#3 FNO-Diffusion hybrid
DICE: 57.03%
DICE by Model
Why the Hybrid Likely Failed
This result does not mean FNO is useless for vision. It suggests that FNO was asked to replace too much of the local segmentation machinery.
What FNO is good at
FNO layers learn global spatial operators by mixing low-frequency Fourier modes. That is powerful when broad structure or resolution-robust mapping matters.
What tumor masks need
Brain tumor segmentation is dense prediction: tiny enhancing regions, irregular edges, and sharp class transitions matter at the pixel level.
What U-shape preserves
Encoder-decoder paths, convolutions, and skip connections preserve local detail while still building coarse semantic context.
What the hybrid lost
Replacing too much of the U-shape pathway with spectral blocks likely reduced boundary refinement and made timestep conditioning harder to stabilize.
Visual Evidence
The qualitative examples make the same point as the DICE table: the model needs mask detail, not only global shape.
FNO Baseline (Figure 4, PDF page 8)
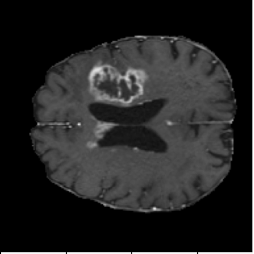


DICE 65.13% · Captures global context but loses boundary precision.
Diffusion with U-shape (Figure 5, PDF page 8)
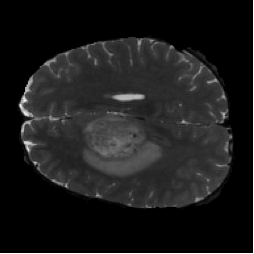

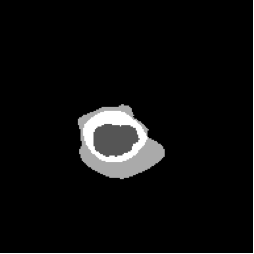
DICE 69.44% · Best qualitative and quantitative segmentation quality.
FNO-Diffusion Hybrid (Figure 6, PDF page 8)
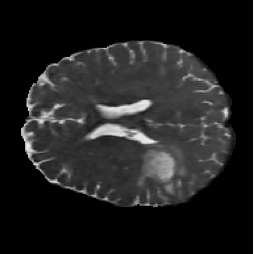

DICE 57.03% · Hybrid underperformed, especially on fine local boundaries.
Why FNO Still Matters
The failed segmentation hybrid does not close the door on FNO. It clarifies where FNO should sit inside a vision architecture.
Classification and global structure
FNO-style models have been explored for image classification, especially where resolution robustness or global frequency structure is useful.
Resolution-invariant image classificationFusion with modern blocks
Like MLP or attention modules, FNO may be most useful as one branch inside a hybrid model: global spectral context plus local detail paths.
Multi-sized image classification with FNODense prediction needs restraint
For segmentation, the design should not ask FNO to carry the full boundary-refinement burden. It should complement, not replace, local modules.
FNO for low-quality image recognitionNext Design I Would Try
The next version should preserve the U-shape backbone and add FNO more carefully.
1. Keep the local U-shape path
Do not remove skip connections and convolutional refinement. They are the strongest inductive bias for boundary precision.
2. Add FNO as a global branch
Use spectral features alongside the local path, then fuse features with attention, gating, or lightweight MLP mixing.
3. Run controlled ablations
Compare local-only, spectral-only, and fused variants while keeping data split, loss, and training schedule fixed.
4. Stabilize before scaling
Stay with 2D slices until the fusion path is stable, then consider 3D extension and stronger multiclass training.
Implementation Lessons
MedSegDiff-V2 was originally the planned foundation for the hybrid model. The goal was to combine FNO with MedSegDiff so the diffusion pipeline could use spectral global context for brain MRI segmentation. In practice, MedSegDiff-V2 was difficult to deploy, and its architecture did not provide a clean insertion point for FNO blocks. Adding FNO looked less like a modular extension and more like changing the whole model structure, so the experiment shifted toward a simpler diffusion U-shape baseline where each component could be isolated and tested.
What became workable
- The diffusion U-shape pipeline offered a clearer modular structure than MedSegDiff-V2.
- Forward diffusion, denoising, and loss components were easier to isolate for debugging.
- Swapping components, including FNO blocks, was straightforward in the modular baseline.
What stayed difficult
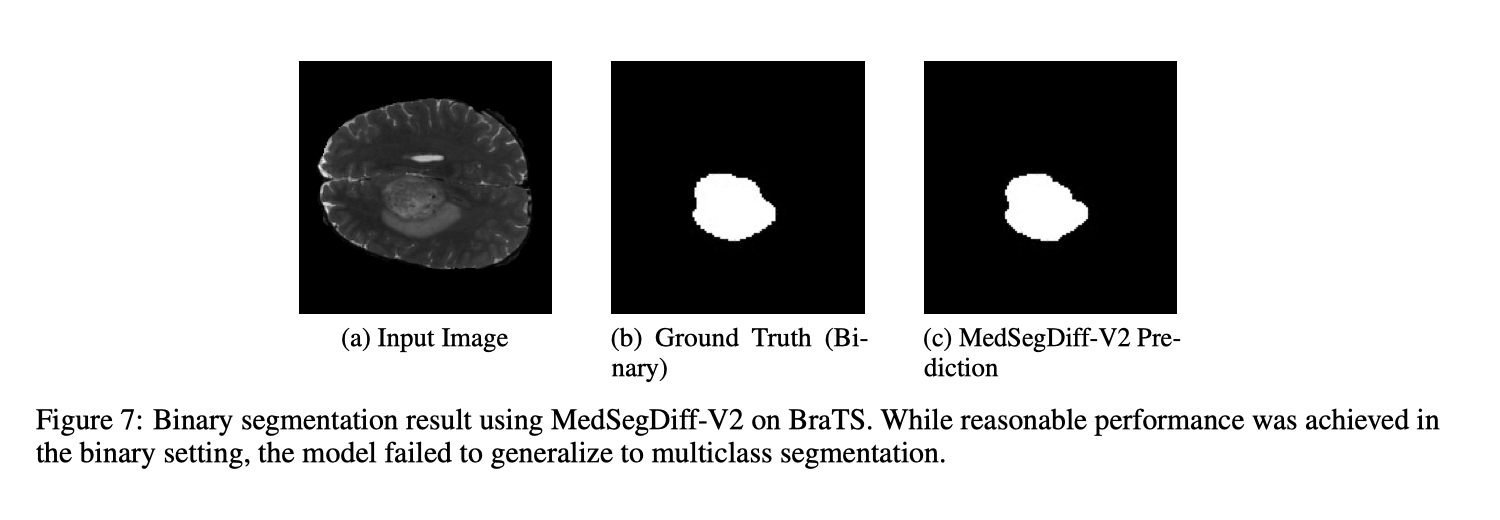
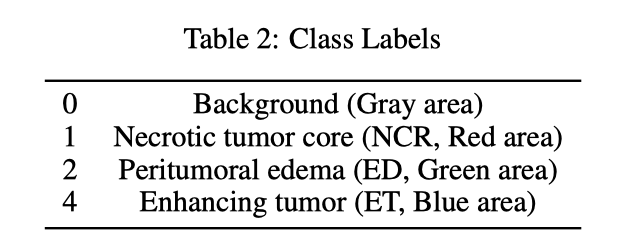
- Reproducing MedSegDiff-V2 for multiclass segmentation failed despite reasonable binary performance.
- Deployment and training were unstable, making multiclass tuning hard to trust.
- Timestep embeddings with FNO blocks caused shape mismatches and conditioning issues.